Scientists use super-resolution microscopy to study previously undiscovered cellular worlds. It allows them, for example, to examine subcellular architecture and molecular interactions at the nanoscale. These methods revolutionised light microscopy and earned its inventors the 2014 Nobel Prize in Chemistry.
To understand the impact of this technology, let us first consider the limits of previously existing techniques. Fluorescence microscopy measures the light emitted by so-called fluorophores that are attached to biological structures of interest. Like in other forms of light microscopy, the resolution that can be achieved this way is limited by the physical properties of light itself. When looking at tiny objects, for example a fluorescently labeled molecule, what we observe is a bright spot which is much bigger than the molecule itself. If two such light sources are too close to each other, their images blur into each other and they can no longer be distinguished from each other.
Super-resolution microscopy methods overcome this fundamental limit. One of these methods is single-molecule localization microscopy which circumvents the resolution limit with a simple trick: Instead of activating all fluorophores in the specimen at the same time, only a few fluorophores are randomly activated at each time point, resulting in a series of images with isolated spots that can be localized with high precision. Subsequently, localizations from many such images are combined to reconstruct a super-resolved image.
However, this method has one major drawback: typically, achieving a high localization precision requires the spots to be well separated. Therefore, prior to our development, one needed to make sure that only a few spots were visible in each image, which in turn resulted in long imaging times.
The DECODE algorithm uses deep learning to enable precise localisation under difficult conditions
In a collaborative effort between the labs of Jakob Macke (University of Tübingen), Jonas Ries (EMBL Heidelberg) and Srinivas Turaga (Janelia Research Campus, Virginia, USA), led by Artur Speiser and Lucas-Raphael Müller, we developed an algorithm called DECODE that overcomes this limitation. It accurately detects and localizes fluorophores at high activation densities, which enables up to ten times higher imaging speeds with minimal loss of resolution. Our results have been published in Nature Methods.
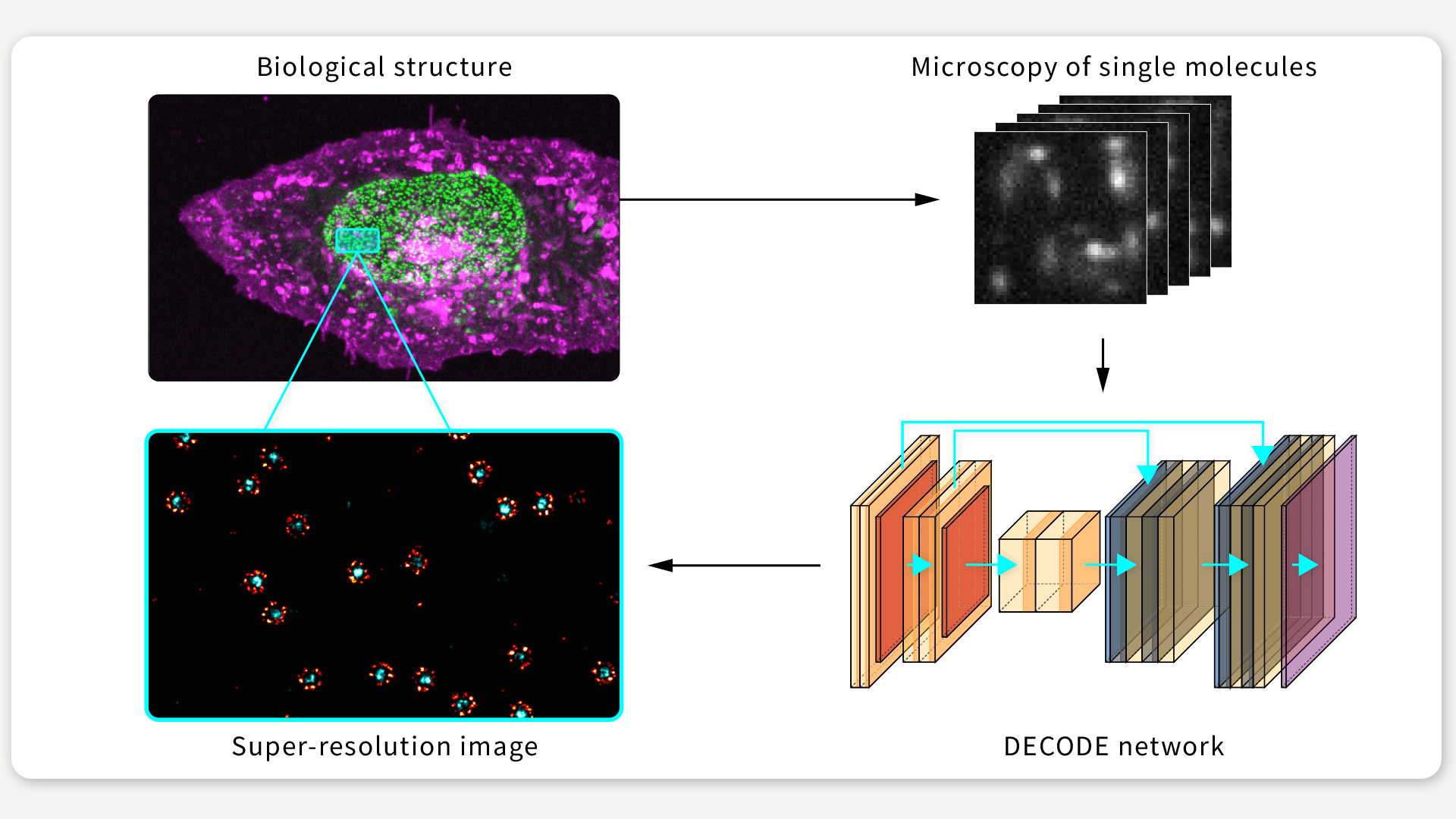
FIGURE 1 / DECODE is a novel algorithm that increases the performance in single-molecule localization microscopy experiments. Figure: Jonas Ries
Our algorithm is based on deep learning: We use an artificial neural network which takes camera images as input, and produces predictions of the underlying fluorophore locations (See Figure 1). To make this work, we had to solve several challenges.
Training neural networks requires pairs of inputs and reference labels, i.e. the desired network outputs, in our case the locations of the fluorophores. However, in our application, we do not know where exactly the fluorophores are. We therefore used a method that has recently become popular for training neural networks in such situations, and which the group has also successfully used in other contexts. Rather than training the network on real images, we used synthetic data that we generated by numerical simulation.
By carefully incorporating what we know about the microscope, the camera, and the molecules’ patterns of fluorescence, we achieve a close similarity between real data and simulations. As a result, networks trained to localize fluorophores on simulated data are then also able to do it on real images!
DECODE allows for ten times higher imaging speeds
Furthermore, we developed a network architecture and loss function that enables our network to localize fluorophores with a precision that is close to the theoretical limit. At the same time it produces uncertainty estimates, i.e. it tells us about the quality of each localization. Removing poor localizations can further enhance the final resolution of the image.
As Figure 2 illustrates, we were able to show that DECODE maintains a high resolution, even when working on emitter densities that are around 10 times higher than those usually employed in single-molecule localization microscopy.
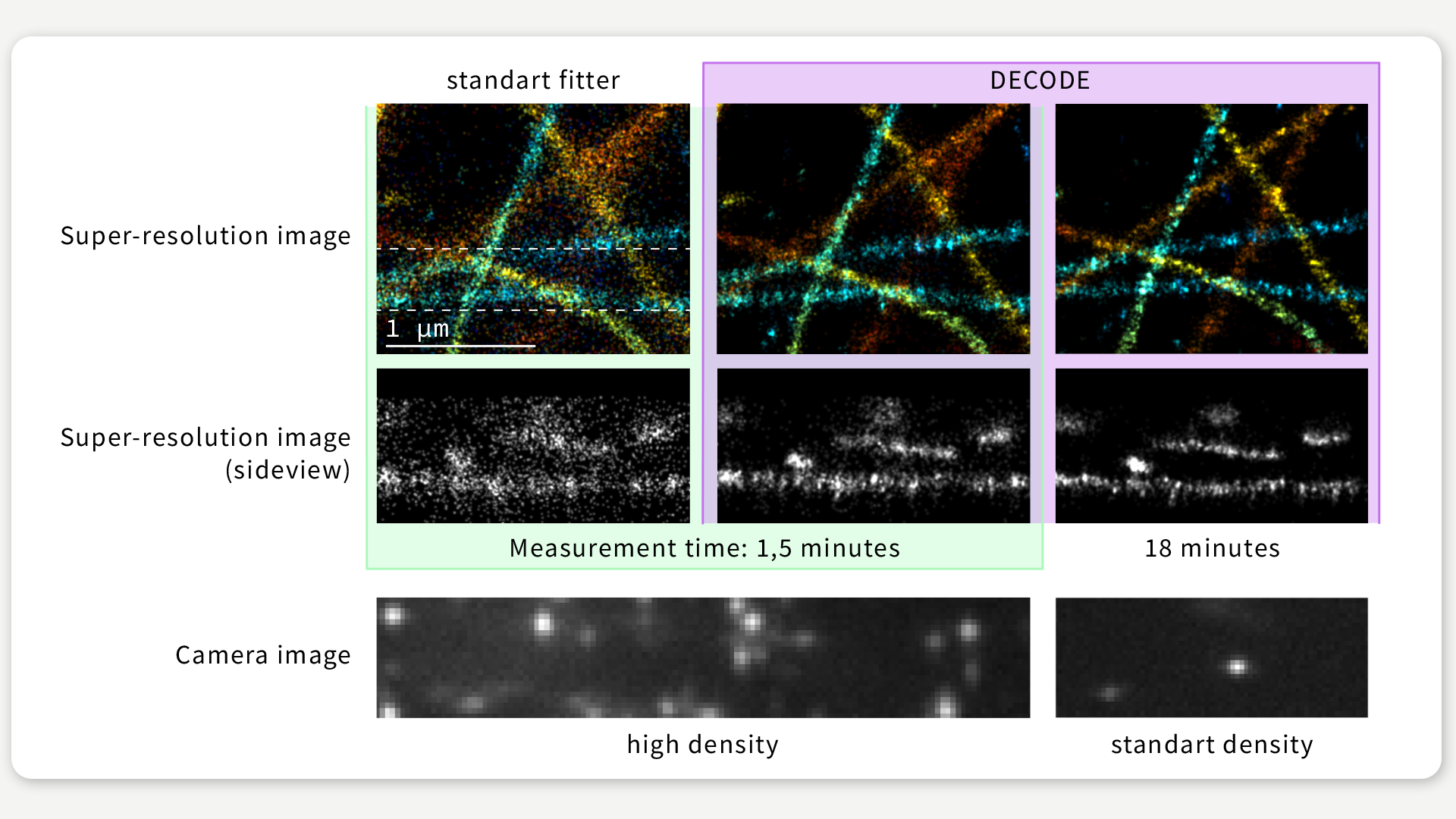
FIGURE 2 / We imaged the same biological structure under different conditions, resulting in higher activation densities and in turn in lower imaging times. Even when imaging with 12 times higher speed the resolution achieved by DECODE only slightly deteriorates, while CSpline, another popular single-molecule localization microscopy algorithm optimized for high-density data, produces a far less detailed reconstruction. Figure: Jonas Ries
Remarkably, on a public challenge that uses authentic simulated datasets to benchmark single-molecule localization microscopy software, DECODE outperformed all 39 algorithms in all conditions, often by a substantial margin. This means that DECODE is always the algorithm of choice, not only for particularly dense data.
With DECODE many labs can improve their images
Our work shows how a novel approach to an ostensibly simple problem like the localization of spots can result in considerable advantages. We built a software package which implements the DECODE algorithm and which is available online. We believe that just by upgrading their analysis software to DECODE many labs performing single-molecule localization microscopy can improve their images, even on their existing datasets. More generally, DECODE shows the power of using numerical simulations to train neural networks to solve challenging analysis problems in the natural sciences.
Original publication: Speiser A., Muller L.-R. et al. Deep learning enables fast and dense single-molecule localization with high accuracy. Nat. Methods. https://doi.org/10.1038/s41592-021-01236-x (2021).
Text: Artur Speiser
Cover image: COS-7 cell imaged with single-molecule localization microscopy. Reconstruction by Artur Speiser et. al., data provided by Wesley Legant et. al.
Identifying Models in
Neuroscience with Machine Learning

Comments